The use of multiomics technologies and analysis for research continues to expand in academia and across industry. Multiomics assays measure multiple sources of biological information from a single sample. In the single-cell context, examples include CITE-seq (RNA gene expression and antibody-based protein expression), 10x Multiome (transcriptomics with chromatin accessibility) and spatial transcriptomics approaches (cellular or subcellular locations along with gene expression). Multiomics analysis aims to extract meaning from multiple modalities, either from a multiomics technology or by combining data from separate assays.
Why multiomics matters
Multiomics technologies can be a powerful tool to learn more about a biological system in a variety of ways. Measuring complementary modalities can provide an orthogonal viewpoint. For example, having gene expression and lipidomics from the same samples can help discover links between parts of the metabolic system that are not visible from a single modality. Analysis can also benefit from additional context, such as spatial locations being used to confirm whether ligand-recepter pairs expressed in different cell types are physically close enough to interact, something that cannot be inferred from dissociated cells alone. Paired multiomics measurements can also be used to validate results, for example confirming that changes in gene expression are correlated with protein expression in the same cells.
The challenges of multiomics analysis
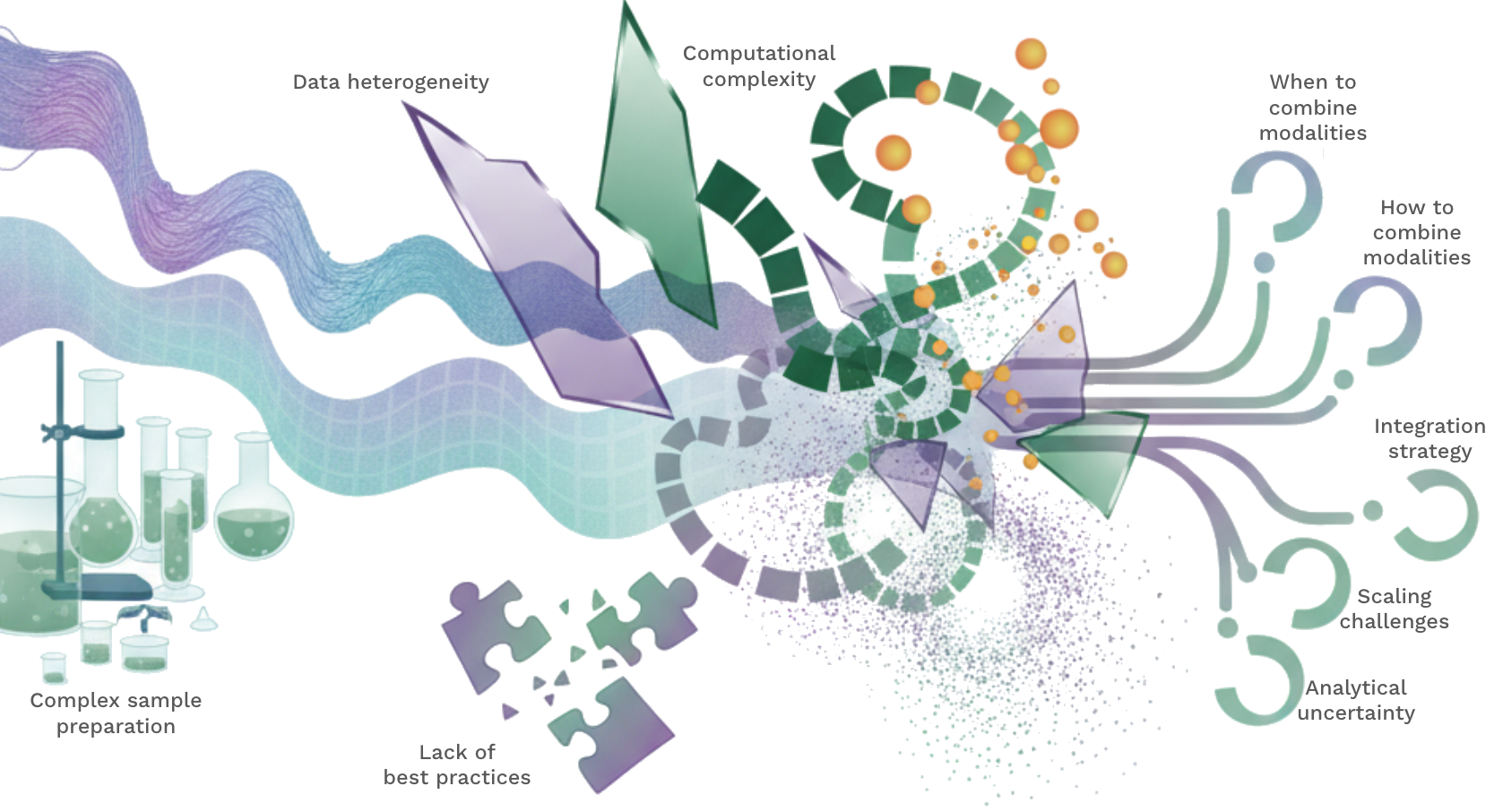

While multiomics opens up a range of possibilities, it also comes with a new set of challenges. Multiomics assays are often more expensive and can require more complex sample preparation steps. Computational analysis of multiomics can be significantly more complex and requires analysts to become familiar with several tool sets and algorithms as each modality has its own processing steps, assumptions and limitations. Less common modalities often do not have established best practices for analysis and require more experimentation with different approaches. Scale also presents a challenge, as does the variety of file types and measured values. When and how to combine modalities is a particularly crucial decision. Depending on the technologies used and the research question, it may make sense to concatenate measurements before analysis, use an integration method to place them in a shared space or to analyse each modality separately and combine the results.
Toward robust and scalable solutions
As the field matures and organisations develop internal practices many of the computational challenges can be addressed by developing standard processing workflows, allowing different approaches to be run in parallel while making use of available compute resources.
Multiomics is not just about adding more data layers - it is about building the right analytical foundations to turn that complexity into insight.